Bacteriophages, or phages, are a type of virus that specifically target bacteria for resources and reproduction1. The diversity of phages in size, functional capability, and genetic information is exponentially increasing as more are discovered and characterized. In general, phage infect a specific species of bacteria. Almost all phages have similar structures, consisting of a capsid “head” that stores genetic information and a “tail” that is used to interact with a host. Like other viruses, phages do not have a nucleus. However, it was discovered by Vorrapon Chaikeeratisak et al. in 2017 that a set of phages that specifically infect Pseudomonas species were observed with a proteinaceous, nucleus-like compartment, or “shell”, that surrounded the phage DNA2. An example of this can be seen in Figure 4A from the paper, where they used cryo-electron tomography to show an infected cell that contained phage with the shell, represented by the phage that are darker in color.
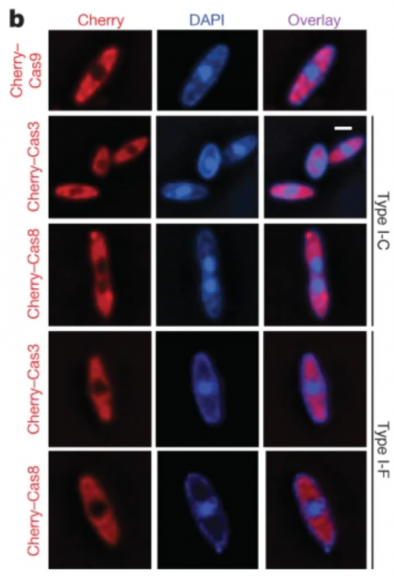
This preliminary finding influenced the work of Senén Mendoza from the Bondy-Denomy group, who normally study CRISPR-Cas biology and function. In 2020, Mendoza and others published a paper entitled “A bacteriophage nucleus-like compartment shields DNA from CRISPR nucleases”, where they find that the previously studied “shell” can actually protect phage DNA from degradation by host CRISPR machinery3. In this study, they found that DNA the large phage ΦKZ, which exclusively infects Pseudomonas aeruginosa and was part of the earlier group of phages studied by Chaikeeratisak, was unaffected when the phage was challenged with an array of CRISPR enzymes. Some phages that infect P. aeruginosa have been previously shown to block CRISPR enzymes through the production of inhibitory proteins. However, the authors performed a genome-wide search and discovered that ΦKZ did not encode for any of these proteins. Through the use of fluorescence microscopy, they were able to show that different CRISPR-Cas enzymes are unable to breach the “shell”, as shown here in Figure 2b from the paper. Here, the first column shows CRISPR enzymes labeled with mCherry and the second column shows phage DNA stained with DAPI. They also explain that, when removed from the “shell”, ΦKZ phage DNA is able to be cleaved by CRISPR enzymes, indicating that the lack of degradation is not due to an intrinsic feature of the DNA. Finally, the authors created a fusion protein of different CRISPR enzymes to a host protein involved in DNA replication that had been known to bypass the “shell”, though the specific mechanism by which this is achieved is unknown. This resulted in cleavage of ΦKZ DNA, again indicating that the nucleus-like compartment is what confers resistance to CRISPR enzymes.
The authors conclude that this discovery speaks to the larger trend of growing phage diversity and highlights a unique and intriguing point of evolution in the viral realm. They propose that this shell could be observed in other, large phages that are not specific to Pseudomonas sp. One area in which the authors plan to continue researching is understanding how the “shell” selectively permits which enzymes are able to reach the phage DNA. This paper represents one of a growing number of reports that highlight unique features of viruses that defy expectations.
1. Kasman LM, Porter LD. Bacteriophages. [Updated 2020 Oct 1]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2020 Jan-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK493185/
2. Mendoza, Senén D., et al. "A bacteriophage nucleus-like compartment shields DNA from CRISPR nucleases." Nature 577.7789 (2020): 244-248.
3. Chaikeeratisak, Vorrapon, et al. "Assembly of a nucleus-like structure during viral replication in bacteria." Science 355.6321 (2017): 194-197.